PAF1 transcriptional regulatory sub-network
The possible genes regulated by the transcription factor were obtained by calculation, and the results were displayed in the form of network
NETWORK ID: CN1181
In order to ensure the specificity of data, an ID index is built by the database itself, and each result corresponds to an ID.
CONDITION: mestanolone
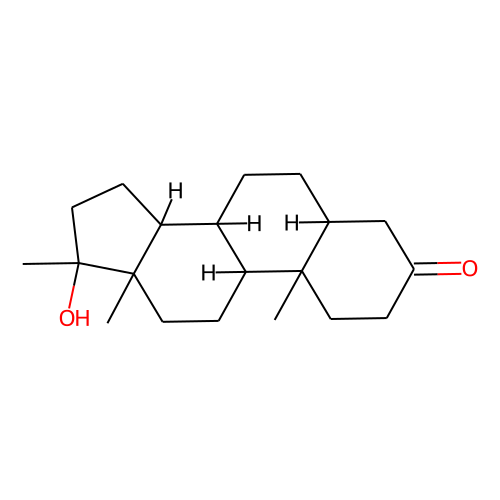
| ID | BRD-K18523449 |
| name | mestanolone |
| type | Small molecule |
| molecular formula | C20H32O2 |
| molecular weight | 304.5 |
| condition dose | 10 uM |
| condition time | 24 h |
| smiles | C[C@]12CCC(=O)C[C@@H]1CC[C@@H]3[C@@H]2CC[C@]4([C@H]3CC[C@]4(C)O)C |
CONDITION TYPE: trt_cp
Compound
CELL LINE: MCF7
Cell lines used for experiments
TRANSCRIPTION FACTOR: PAF1
Transcription regulatory factors corresponding to this network
Differential expressed genes in this transcriptional regulatory sub-network
| logfc | > 0.45 and P < = 1e-5. If you want to view or download the data, you can click [expand]
| Symbol | control | control | control | condition | condition |
|---|---|---|---|---|---|
| HMOX1 | 5.2332 | 5.1734 | 5.1631 | 3.376 | 4.0403 |
| SUV39H1 | 7.8992 | 7.836 | 8.6959 | 4.8763 | 6.4013 |
| WIPF2 | 7.1944 | 6.8491 | 6.7711 | 5.2447 | 5.7002 |
| EML3 | 8.0266 | 7.7962 | 8.1981 | 6.185 | 6.5949 |
| PXN | 11.7483 | 11.9648 | 11.7304 | 10.2849 | 10.6294 |
| ZNF274 | 7.4086 | 7.1197 | 7.1944 | 5.7458 | 5.9685 |
| VAPB | 9.5134 | 8.9772 | 9.0941 | 6.6539 | 7.8862 |
| SCARB1 | 8.8931 | 8.7387 | 9.7807 | 6.3679 | 7.5753 |
| LSM6 | 8.069 | 7.7315 | 8.2688 | 5.9511 | 6.7709 |
| SACM1L | 8.4684 | 8.4858 | 7.4934 | 6.0596 | 6.5202 |
| ABCF3 | 9.3399 | 8.6539 | 9.8097 | 7.0911 | 7.482 |
| RAD9A | 7.2572 | 7.2223 | 7.2361 | 6.0596 | 5.78 |
| TSTA3 | 6.7189 | 6.7151 | 6.8592 | 5.2447 | 5.4966 |
| ABCB6 | 8.8156 | 8.4545 | 9.2476 | 5.9511 | 6.7866 |
| CIRBP | 10.7165 | 9.6743 | 10.8593 | 8.3645 | 8.6701 |
| CRTAP | 6.2176 | 6.656 | 7.0675 | 4.6429 | 5.2698 |
| ANXA7 | 9.6723 | 9.4022 | 9.5316 | 7.6978 | 7.9076 |
| SKIV2L | 7.7985 | 7.5049 | 8.1616 | 6.2785 | 6.4269 |
| ATP1B1 | 8.8784 | 8.5755 | 9.9191 | 5.9511 | 7.3499 |
| ADO | 6.6813 | 6.3485 | 6.1977 | 4.6429 | 5.1556 |
| RNH1 | 10.8062 | 10.6559 | 10.6416 | 9.5389 | 9.5282 |
| NOSIP | 8.6348 | 8.5284 | 9.1542 | 7.1353 | 7.5479 |
| ZMYM2 | 10.7206 | 10.5178 | 10.468 | 8.3645 | 9.4507 |
| GRN | 10.1787 | 10.7544 | 11.0535 | 6.5792 | 8.159 |
| TLE1 | 9.4202 | 9.2338 | 9.8901 | 7.7677 | 8.2026 |
| ACOT9 | 8.4342 | 7.764 | 8.3743 | 5.4711 | 5.9857 |
| F12 | 10.2752 | 10.6456 | 10.2965 | 8.0355 | 9.1524 |
| CAPNS1 | 10.3182 | 9.981 | 10.8824 | 8.6327 | 7.7434 |
| SF3B2 | 9.2006 | 8.728 | 9.882 | 6.912 | 7.7516 |
| MATR3 | 9.7884 | 9.8635 | 9.9275 | 8.3696 | 8.8674 |
| CD81 | 12.9719 | 12.6233 | 12.9921 | 10.4738 | 10.3851 |
| S100A10 | 11.9304 | 12.0301 | 11.9165 | 6.9518 | 9.905 |
| TRIM28 | 11.3335 | 10.5696 | 11.9066 | 8.3515 | 9.2542 |
| STOM | 8.4467 | 7.0786 | 7.8178 | 5.4793 | 5.5813 |
| CYC1 | 11.7322 | 12.0988 | 12.1574 | 8.9 | 10.5746 |
| SYNGR2 | 12.4215 | 12.608 | 11.7084 | 10.4837 | 10.423 |
| HDAC1 | 10.6537 | 11.2178 | 10.5416 | 9.335 | 9.4343 |
| ERP29 | 9.4996 | 10.2852 | 11.1968 | 6.944 | 8.2472 |
| TALDO1 | 9.9783 | 9.2887 | 10.0382 | 8.0953 | 7.9629 |
| NQO1 | 11.4948 | 11.0589 | 12.2017 | 9.5251 | 9.504 |
| WSB2 | 9.9946 | 8.7973 | 9.5206 | 6.2328 | 7.7937 |
| CYB5R3 | 10.4852 | 10.2324 | 10.8831 | 7.398 | 9.1013 |
| CD55 | 7.8509 | 8.1563 | 7.4024 | 5.9338 | 5.9517 |
| CIB1 | 9.8454 | 9.1988 | 9.5886 | 5.395 | 7.0332 |
| ARPC1B | 10.2838 | 10.6253 | 11.1221 | 7.3031 | 8.5825 |
| CHMP2A | 9.3966 | 9.1212 | 8.8505 | 5.0161 | 7.1814 |
| RAB13 | 12.0968 | 12.6372 | 12.2369 | 11.119 | 10.4767 |
| CTSH | 9.2133 | 7.9626 | 8.6869 | 5.58 | 4.7767 |
| EPS8 | 11.2991 | 10.3003 | 12.2136 | 6.9721 | 6.8028 |
| CTPS1 | 7.9272 | 8.2325 | 9.0801 | 6.0702 | 6.892 |
| SNRPD1 | 12.0427 | 11.7639 | 12.0689 | 10.7522 | 10.7979 |
| LSM4 | 11.1024 | 10.5005 | 11.1618 | 7.1822 | 9.2627 |
| UBE2S | 11.6748 | 11.9189 | 13.067 | 5.7958 | 8.9773 |
| GCC2 | 6.9878 | 6.3332 | 6.3335 | 4.3968 | 5.2705 |
| ZMPSTE24 | 11.0936 | 11.1732 | 11.3617 | 8.4657 | 9.6061 |
| SLPI | 4.9879 | 5.1446 | 4.9761 | 3.4042 | 3.2206 |
| BMS1 | 9.4147 | 9.1404 | 9.1808 | 7.2555 | 8.1459 |
| TBC1D4 | 6.2538 | 4.0988 | 5.1439 | 0.0269 | 2.4934 |
| PROCR | 11.6217 | 11.4706 | 11.5767 | 9.624 | 10.5095 |
| COIL | 10.2724 | 9.9727 | 10.1917 | 7.9888 | 8.843 |
| EMP3 | 5.2494 | 5.535 | 6.3528 | 1.8852 | 2.3429 |
| GCLM | 8.0953 | 7.0158 | 7.9053 | 5.1665 | 5.9064 |
| WWP2 | 7.9338 | 8.0033 | 8.4545 | 5.392 | 6.8092 |
| PCBP2 | 13.8135 | 13.615 | 14.2581 | 11.2936 | 12.5107 |
| TSC22D2 | 8.8333 | 8.7628 | 7.8491 | 5.8152 | 6.7702 |
| TFF1 | 12.3368 | 12.8006 | 12.9943 | 6.0523 | 9.0827 |
| ARL4A | 4.2839 | 3.8908 | 4.6177 | 2.5597 | 3.0634 |
| PSCA | 6.6337 | 6.428 | 7.3804 | 3.3987 | 4.747 |
| CCDC22 | 9.2025 | 8.9775 | 9.2556 | 7.3458 | 7.9546 |
| GRB14 | 7.8167 | 7.0214 | 7.517 | 4.84 | 5.3659 |
| NDRG2 | 7.5006 | 8.4124 | 9.4763 | 4.1658 | 6.188 |
| KAT5 | 7.2693 | 7.8778 | 7.6932 | 5.9881 | 6.3769 |
| RPL36AL | 12.5616 | 12.2471 | 12.8219 | 10.7348 | 10.2182 |
| ATP6AP1 | 8.4902 | 8.4588 | 9.376 | 6.2971 | 7.0964 |
| CLIC1 | 10.6189 | 10.5168 | 10.431 | 8.5826 | 8.5543 |
| TAPBP | 8.0035 | 7.7458 | 8.4959 | 6.7163 | 6.3834 |
| NEU1 | 7.7555 | 8.1266 | 7.8671 | 6.6636 | 6.6097 |
| SLC25A11 | 12.9079 | 11.7508 | 11.6797 | 9.4496 | 10.2479 |
| ISCU | 10.258 | 9.189 | 9.8485 | 7.9022 | 7.9906 |
| NR2F2 | 9.3519 | 9.477 | 10.671 | 7.9768 | 6.3034 |
| HIGD2A | 13.0212 | 13.0769 | 13.6518 | 11.7805 | 11.8857 |
| PRPS1 | 8.3865 | 9.4024 | 8.5596 | 6.67 | 7.2773 |
| POP7 | 10.052 | 10.0665 | 10.2973 | 8.7873 | 8.7386 |
| PRKAA1 | 8.198 | 8.4347 | 7.9187 | 6.4212 | 6.793 |
| TNFSF13 | 7.3921 | 6.1832 | 6.1379 | 4.4041 | 4.6848 |
| YWHAE | 13.6008 | 13.6941 | 13.3705 | 11.2559 | 12.1994 |
| SERPINB6 | 10.4209 | 9.6621 | 10.8806 | 6.6069 | 8.0129 |
| BAG1 | 7.8595 | 7.2029 | 7.7302 | 5.8364 | 5.5651 |
| TMPRSS2 | 5.4706 | 5.1678 | 5.0307 | 1.8711 | 3.4992 |
| B3GALNT1 | 7.3672 | 6.5028 | 7.64 | 4.7899 | 5.5695 |
| EIF4B | 9.8153 | 9.1084 | 9.6622 | 6.1828 | 7.3351 |
| AHNAK | 8.6057 | 8.216 | 7.7778 | 5.1769 | 6.1296 |
| PTOV1 | 9.9752 | 9.7922 | 10.3713 | 8.7086 | 8.76 |
| POLDIP3 | 7.2605 | 7.1986 | 7.3016 | 5.6476 | 6.2445 |
| PSME4 | 8.8628 | 8.1562 | 8.8046 | 6.302 | 7.3036 |
| CMTR1 | 7.9204 | 7.9343 | 7.7824 | 6.0308 | 6.6567 |
| MFSD5 | 9.7091 | 9.2832 | 9.3869 | 7.1842 | 8.0382 |
| GNPTAB | 8.5821 | 7.4484 | 7.6187 | 5.7533 | 5.9221 |
| APRT | 12.4435 | 12.462 | 13.2549 | 10.9054 | 10.63 |
| HNRNPA1 | 11.8602 | 12.5616 | 11.9304 | 9.9747 | 10.8243 |
| BRD2 | 6.5958 | 6.7623 | 7.1664 | 5.5811 | 4.7515 |
| SEC16A | 7.0521 | 7.3509 | 7.3849 | 3.8827 | 5.7045 |
| TRIOBP | 8.5126 | 8.1601 | 8.2828 | 5.9856 | 7.0478 |
| HLA-E | 6.3546 | 6.5184 | 6.348 | 4.7794 | 4.8436 |
| COPZ1 | 8.7393 | 8.9239 | 9.3832 | 7.4953 | 7.6471 |
| NUCKS1 | 12.5487 | 12.0667 | 13.615 | 10.0367 | 10.4013 |
| GOLPH3 | 9.4189 | 9.4294 | 9.3503 | 7.4253 | 7.969 |
| EMC3 | 10.159 | 9.7354 | 9.841 | 6.7987 | 8.4559 |
| AKIRIN1 | 7.5954 | 7.2114 | 7.5706 | 5.3428 | 6.0261 |
| SPCS1 | 9.4402 | 8.5947 | 9.1558 | 7.2549 | 7.4979 |
| VPS51 | 9.5147 | 9.3515 | 9.7621 | 7.8003 | 8.0119 |
| PRC1 | 9.8427 | 9.8104 | 9.3797 | 6.6414 | 8.2199 |
| TSEN34 | 11.0112 | 11.1129 | 10.9702 | 8.8148 | 9.2352 |
| MLPH | 12.1509 | 11.999 | 11.9256 | 7.0567 | 9.8835 |
| RMND5B | 9.9661 | 9.515 | 9.7574 | 7.9028 | 8.5851 |
| SPATS2 | 8.6999 | 8.5685 | 8.534 | 7.4007 | 7.0393 |
| U2AF2 | 10.0523 | 9.9661 | 9.6222 | 7.5511 | 8.0574 |
| SFXN1 | 9.9911 | 9.751 | 10.8568 | 8.3722 | 8.4131 |
| MRPS30 | 9.1948 | 8.3029 | 9.1364 | 4.9151 | 6.1381 |
| THEM6 | 8.7968 | 8.7902 | 8.7581 | 6.842 | 7.6726 |
| NDUFA3 | 11.2374 | 10.674 | 10.6585 | 8.4425 | 8.4321 |
| ZDHHC7 | 8.7664 | 7.8728 | 7.9843 | 5.5728 | 6.7766 |
| SLC25A15 | 8.093 | 7.941 | 8.313 | 5.4397 | 6.5454 |
| LXN | 15 | 14.9004 | 13.7202 | 12.3268 | 12.1541 |
| BRF2 | 7.3982 | 7.1491 | 7.6706 | 5.4281 | 6.1461 |
| WWOX | 6.9808 | 6.3867 | 6.8975 | 5.1795 | 5.0418 |
| MACROD1 | 9.5381 | 9.5208 | 10.3534 | 6.9948 | 8.2841 |
| HSPA14 | 9.2618 | 8.5457 | 8.3674 | 7.2047 | 7.0735 |
| SLC6A14 | 8.1835 | 6.4506 | 8.1042 | 4.6093 | 5.2406 |
| RBM23 | 8.8224 | 8.9556 | 8.976 | 6.7648 | 7.6851 |
| THADA | 7.2665 | 6.9643 | 7.1558 | 5.7316 | 5.939 |
| ZNF580 | 10.9815 | 10.985 | 10.9746 | 9.4196 | 9.8936 |
| KLHDC4 | 8.5746 | 8.3043 | 8.3291 | 7.0942 | 7.2917 |
| SF3B5 | 10.9047 | 10.8101 | 10.7913 | 9.1386 | 9.8451 |
| IMP3 | 9.3575 | 8.9841 | 9.7265 | 7.3455 | 8.1142 |
| UGCG | 6.059 | 6.4411 | 7.0657 | 3.8224 | 4.5044 |
| KANSL2 | 8.4855 | 8.6499 | 8.4429 | 6.4875 | 6.9357 |
| CSPG5 | 7.1184 | 6.6882 | 7.2429 | 5.237 | 5.3433 |
| SSH3 | 9.697 | 9.7099 | 10.3812 | 8.2971 | 8.4904 |
| ZMIZ2 | 6.1383 | 5.6013 | 6.503 | 4.7389 | 3.965 |
| CORO1B | 11.4721 | 11.5952 | 11.6396 | 9.0559 | 9.4652 |
| CYP1A2 | 6.1682 | 6.5467 | 7.22 | 4.3968 | 5.2222 |
PAF1 transcriptional regulatory sub-network
The possible genes regulated by the transcription factor were obtained by calculation, and the results were displayed in the form of network
ARTICLE: PAF1
[1] PD2/PAF1 at the Crossroads of the Cancer Network.
[2] PAF1 regulation of promoter-proximal pause release via enhancer activation.
[3] Emerging Insights into the Roles of the Paf1 Complex in Gene Regulation.
[5] Cigarette Smoke Induces Stem Cell Features of Pancreatic Cancer Cells via PAF1.
[7] PAF1, a Molecular Regulator of Promoter-Proximal Pausing by RNA Polymerase II.
[10] The Paf1 Complex Broadly Impacts the Transcriptome of Saccharomyces cerevisiae.